Jump to: Ongoing Research Projects
Epigenetic modifications regulate gene expression and allow for tissue-specific expression of transcripts during development and in differentiated cells.
Cancer cells often have markedly different “epigenomes” than normal cells and exhibit profound changes in DNA methylation (a silencing epigenetic mark) of cytosines at CpG dinucleotides. These changes include global loss of methylation at regions such as repetitive elements that must be silenced for genome stability and gain of methylation at the promoter regions of tumor suppressor and other genes. DNA methyltransferase inhibitors (DNMTis) cause re-expression of genes that are silenced by promoter DNA methylation, reactivating tumor suppressor genes.
Transient exposure of multiple types of tumor cells to low doses of DNMTis promotes induction of apoptosis, reduced cell cycle activity, and decreased stem cell functions in cancer cells. Clinical efficacy of DNMTis such as 5-azacytidine (Aza) and 5-aza-2’-deoxycytidine has led to FDA approval for the pre-leukemic disorder myelodysplasia (MDS). Our work has shown that epigenetic therapy with DNMTis “boosts” immune signaling from tumors through activation of the interferon response by double-stranded RNA including hypermethylated endogenous retroviruses (ERVs). We have shown that a major effect of DNMTis is upregulation of immune signaling across solid tumors (breast, colon, and ovarian cancers), which holds true in patient samples from DNMTi clinical trials (Li, Chiappinell et al., Oncotarget 2014)
A key component of this dsRNA is endogenous retrovirus transcripts (ERVs), normally hypermethylated, which become demethylated and overexpressed upon DNMTi treatment. Working with Pamela Strissel and Reiner Strick at the University of Erlangen, we linked upregulation of hypermethylated ERVs to the DNMTi-induced immune response. Inhibiting the interferon response rescues about half of the apoptosis induced by 5-azacytidine, and that 5-azacytidine sensitizes a preclinical model of melanoma to anti-CTLA-4 (immune) therapy (Chiappinelli et al., Cell 2015). This epigenetic treatment upregulates interferon signaling in the tumor cells and causes host immune cells to selectively target the tumor cells for destruction.
Ongoing Research Projects
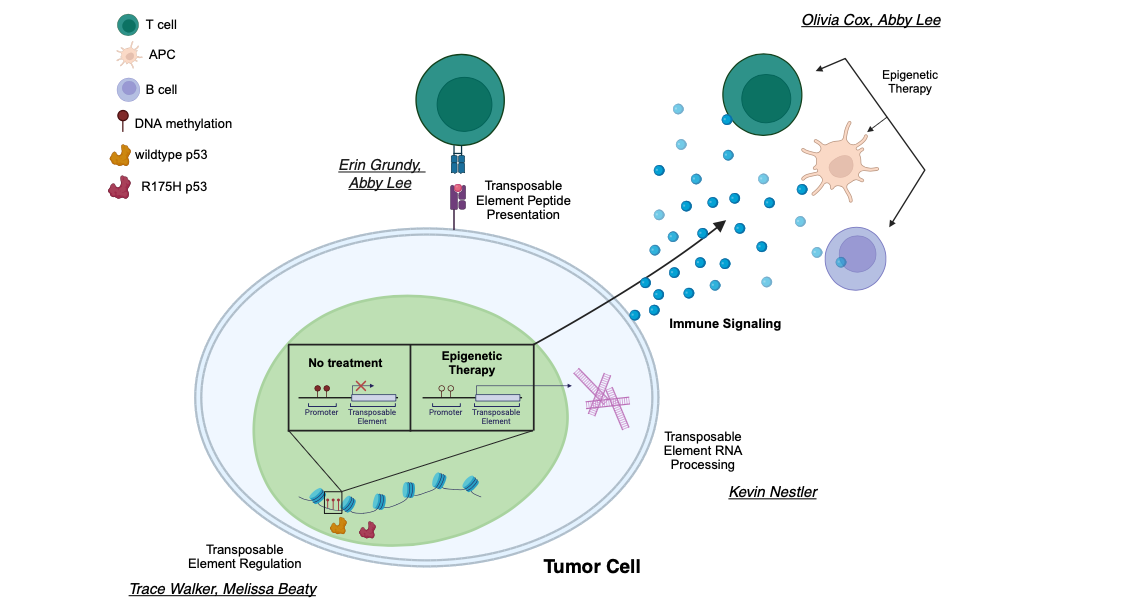
- P53 regulation of transposable element expression and downstream immune signaling in cancer: We seek to understand how transcriptional regulation of TEs by p53 modulates anti-tumor immunity. We showed that wild type p53 binds directly to TEs, activating transcription (learn more). We hypothesize that mutant p53 aberrantly activates TEs, amplifying the TE-induced immune response. Our ongoing work is determining the role of mutant p53 in direct and indirect regulation of TE expression and it's downstream effects.
- RNA modulation of transposable elements in cancer: The enzyme ADAR1 edits immunogenic TE RNA and promotes resistance to DNMTi, specifically in the p53 mutant background. We showed that in a murine model of OC, loss of Adar1 significantly prolongs survival, which is further improved with DNMTi treatment (learn more). We hypothesize that TEs are differentially edited in mutant P53 backgrounds and that disrupting editing activates type I interferon signaling to promote immunologic control of the tumor. Ongoing work is also actively investigating other post-transcriptional modifications of TE RNA and how these affect dsRNA formation and TE immunogenicity.
- Understanding how transposable elements are regulated during cancer progression: Transposable elements are expressed at high levels in embryonic stem cells but are silenced by DNA methylation and histone modifications in most differentiated cells. During the global epigenetic dysregulation that occurs through cancer transformation, TEs lose silencing and become transcribed, and in some cases, active. We showed that TEs lose DNA methylation and gain expression during transformation (learn more). Current work focuses on the immune consequences of TE expression in the early stages of ovarian cancer, which is characterized by high TE expression yet an immune suppressive microenvironment.
- Transposable elements as novel tumor antigens: Some transposable elements can encode proteins that may be processed, presented, and recognized by T cells as tumor antigens (learn more). We are currently investigating which TEs are translated and overexpressed in cancers for novel T cell therapies, potentially in combination with epigenetic therapies that reverse the immunosuppressive tumor microenvironment.